Interactive 3D Force-Directed Edge Bundling
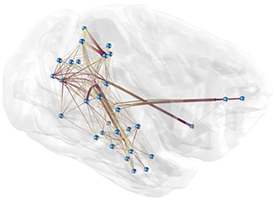
Interactive analysis of 3D relational data is challenging. A common way of representing such data are node-link diagrams as they support analysts in achieving a mental model of the data. However, naïve 3D depictions of complex graphs tend to be visually cluttered, even more than in a 2D layout. This makes graph exploration and data analysis less efficient. This problem can be addressed by edge bundling. We introduce a 3D cluster-based edge bundling algorithm that is inspired by the force-directed edge bundling (FDEB) algorithm [Holten2009] and fulfills the requirements to be embedded in an interactive framework for spatial data analysis. It is parallelized and scales with the size of the graph regarding the runtime. Furthermore, it maintains the edge’s model and thus supports rendering the graph in different structural styles. We demonstrate this with a graph originating from a simulation of the function of a macaque brain.

